Kavi's BGGN 213 Portfolio

Class work for bioinformatics class
Class 11: Structural Bioinformatics pt2
Kavi Gonur (PID: A69046927)
Background
We saw last day that the PDB has 209,886 entries (Oct/Nov 2025). UniProtKB (i.e. protein sequence database) has 199,579,901 entries.
PDB has 0.1051639% coverage of the main sequence database.
209886 / 199579901 * 100
[1] 0.1051639
Enter AlphaFold database (AFDB). https://alphafold.ebi.ac.uk that attempts to provide computed models for all sequences in UniProt.
“AlphaFold DB provides open access to over 200 million protein structure predictions to accelerate scientific research.”
AlphaFold
AlphaFold has 3 main outputs - the predicted coordinates (PDB files) - a local quality score called pLDDT (one for each amino-acid) - a second quality score PAE Predicted Aligned Error (for each pair of amino acid)
We can run AlphaFold ourselves if we not happy with AFDB (i.e. no coverage or poor model).
results_dir <- "HIVPR_dimer_23119.result/HIVPR_dimer_23119/"
pdb_files <- list.files(path=results_dir,
pattern="*.pdb",
full.names = TRUE)
basename(pdb_files)
[1] "HIVPR_dimer_23119_unrelaxed_rank_001_alphafold2_multimer_v3_model_4_seed_000.pdb"
[2] "HIVPR_dimer_23119_unrelaxed_rank_002_alphafold2_multimer_v3_model_1_seed_000.pdb"
[3] "HIVPR_dimer_23119_unrelaxed_rank_003_alphafold2_multimer_v3_model_5_seed_000.pdb"
[4] "HIVPR_dimer_23119_unrelaxed_rank_004_alphafold2_multimer_v3_model_2_seed_000.pdb"
[5] "HIVPR_dimer_23119_unrelaxed_rank_005_alphafold2_multimer_v3_model_3_seed_000.pdb"
library(bio3d)
pdbs <- pdbaln(pdb_files, fit=TRUE, exefile="msa")
Reading PDB files:
HIVPR_dimer_23119.result/HIVPR_dimer_23119/HIVPR_dimer_23119_unrelaxed_rank_001_alphafold2_multimer_v3_model_4_seed_000.pdb
HIVPR_dimer_23119.result/HIVPR_dimer_23119/HIVPR_dimer_23119_unrelaxed_rank_002_alphafold2_multimer_v3_model_1_seed_000.pdb
HIVPR_dimer_23119.result/HIVPR_dimer_23119/HIVPR_dimer_23119_unrelaxed_rank_003_alphafold2_multimer_v3_model_5_seed_000.pdb
HIVPR_dimer_23119.result/HIVPR_dimer_23119/HIVPR_dimer_23119_unrelaxed_rank_004_alphafold2_multimer_v3_model_2_seed_000.pdb
HIVPR_dimer_23119.result/HIVPR_dimer_23119/HIVPR_dimer_23119_unrelaxed_rank_005_alphafold2_multimer_v3_model_3_seed_000.pdb
.....
Extracting sequences
pdb/seq: 1 name: HIVPR_dimer_23119.result/HIVPR_dimer_23119/HIVPR_dimer_23119_unrelaxed_rank_001_alphafold2_multimer_v3_model_4_seed_000.pdb
pdb/seq: 2 name: HIVPR_dimer_23119.result/HIVPR_dimer_23119/HIVPR_dimer_23119_unrelaxed_rank_002_alphafold2_multimer_v3_model_1_seed_000.pdb
pdb/seq: 3 name: HIVPR_dimer_23119.result/HIVPR_dimer_23119/HIVPR_dimer_23119_unrelaxed_rank_003_alphafold2_multimer_v3_model_5_seed_000.pdb
pdb/seq: 4 name: HIVPR_dimer_23119.result/HIVPR_dimer_23119/HIVPR_dimer_23119_unrelaxed_rank_004_alphafold2_multimer_v3_model_2_seed_000.pdb
pdb/seq: 5 name: HIVPR_dimer_23119.result/HIVPR_dimer_23119/HIVPR_dimer_23119_unrelaxed_rank_005_alphafold2_multimer_v3_model_3_seed_000.pdb
rd <- rmsd(pdbs, fit=T)
Warning in rmsd(pdbs, fit = T): No indices provided, using the 198 non NA positions
range(rd)
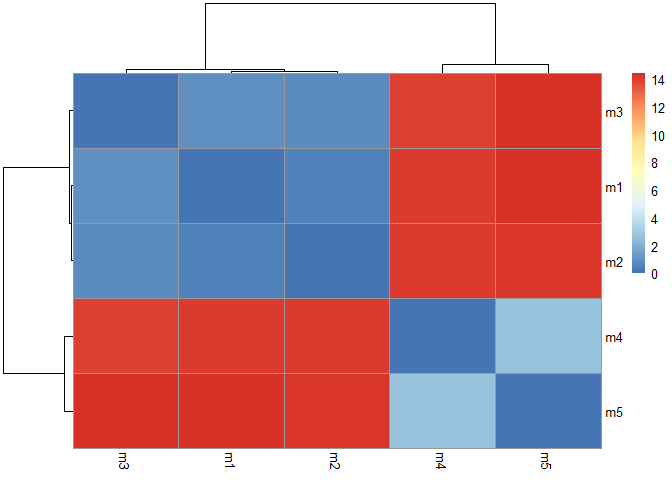
[1] 0.000 14.428
library(pheatmap)
colnames(rd) <- paste0("m",1:5)
rownames(rd) <- paste0("m",1:5)
pheatmap(rd)
pdb <- read.pdb("1hsg")
Note: Accessing on-line PDB file
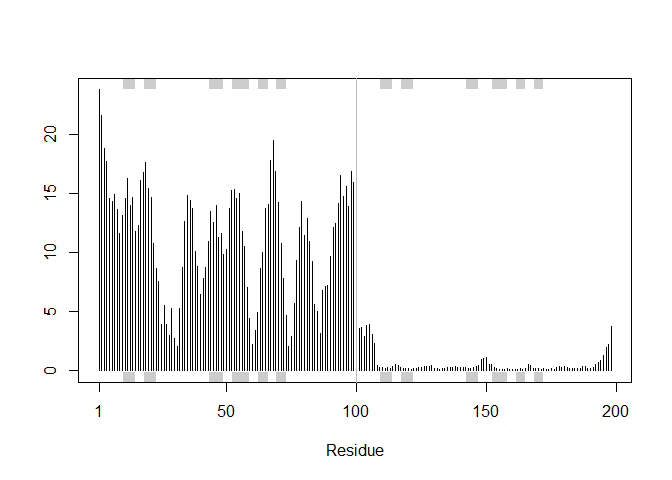
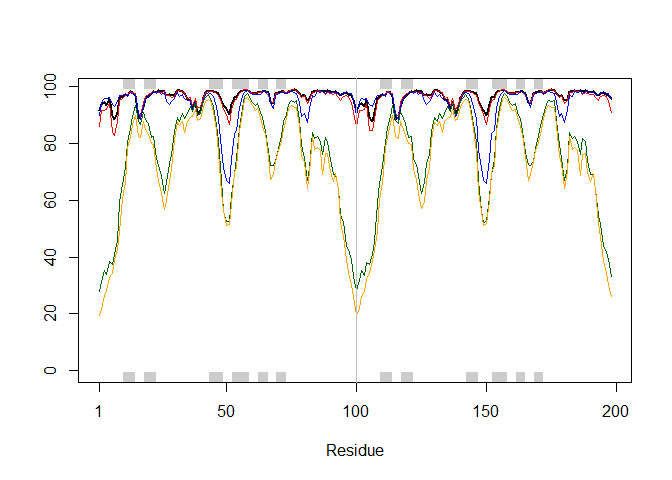
plotb3(pdbs$b[1,], typ="l", lwd=2, sse=pdb)
points(pdbs$b[2,], typ="l", col="red")
points(pdbs$b[3,], typ="l", col="blue")
points(pdbs$b[4,], typ="l", col="darkgreen")
points(pdbs$b[5,], typ="l", col="orange")
abline(v=100, col="gray")
core <- core.find(pdbs)
core size 197 of 198 vol = 8545.079
core size 196 of 198 vol = 7894.973
core size 195 of 198 vol = 3576.564
core size 194 of 198 vol = 1851.077
core size 193 of 198 vol = 1697.205
core size 192 of 198 vol = 1612.775
core size 191 of 198 vol = 1530.208
core size 190 of 198 vol = 1447.408
core size 189 of 198 vol = 1377.117
core size 188 of 198 vol = 1303.826
core size 187 of 198 vol = 1239.04
core size 186 of 198 vol = 1188.139
core size 185 of 198 vol = 1118.473
core size 184 of 198 vol = 1071.674
core size 183 of 198 vol = 1034.016
core size 182 of 198 vol = 980.836
core size 181 of 198 vol = 942.246
core size 180 of 198 vol = 911.395
core size 179 of 198 vol = 879.756
core size 178 of 198 vol = 834.466
core size 177 of 198 vol = 785.268
core size 176 of 198 vol = 762.122
core size 175 of 198 vol = 722.029
core size 174 of 198 vol = 700.395
core size 173 of 198 vol = 677.257
core size 172 of 198 vol = 657.81
core size 171 of 198 vol = 632.907
core size 170 of 198 vol = 614.198
core size 169 of 198 vol = 591.516
core size 168 of 198 vol = 573.979
core size 167 of 198 vol = 552.403
core size 166 of 198 vol = 529.489
core size 165 of 198 vol = 500.545
core size 164 of 198 vol = 482.517
core size 163 of 198 vol = 458.426
core size 162 of 198 vol = 444.455
core size 161 of 198 vol = 433.581
core size 160 of 198 vol = 419.086
core size 159 of 198 vol = 404.934
core size 158 of 198 vol = 393.803
core size 157 of 198 vol = 383.003
core size 156 of 198 vol = 366.654
core size 155 of 198 vol = 352.026
core size 154 of 198 vol = 335.663
core size 153 of 198 vol = 319.398
core size 152 of 198 vol = 307.935
core size 151 of 198 vol = 296.818
core size 150 of 198 vol = 284.289
core size 149 of 198 vol = 273.459
core size 148 of 198 vol = 261.978
core size 147 of 198 vol = 249.6
core size 146 of 198 vol = 237.954
core size 145 of 198 vol = 226.1
core size 144 of 198 vol = 213.265
core size 143 of 198 vol = 200.214
core size 142 of 198 vol = 187.504
core size 141 of 198 vol = 177.525
core size 140 of 198 vol = 167.372
core size 139 of 198 vol = 160.875
core size 138 of 198 vol = 154.455
core size 137 of 198 vol = 148.439
core size 136 of 198 vol = 142.13
core size 135 of 198 vol = 136.529
core size 134 of 198 vol = 130.77
core size 133 of 198 vol = 123.868
core size 132 of 198 vol = 117.609
core size 131 of 198 vol = 112.71
core size 130 of 198 vol = 106.361
core size 129 of 198 vol = 100.591
core size 128 of 198 vol = 95.718
core size 127 of 198 vol = 91.068
core size 126 of 198 vol = 86.862
core size 125 of 198 vol = 82.309
core size 124 of 198 vol = 78.554
core size 123 of 198 vol = 74.632
core size 122 of 198 vol = 70.489
core size 121 of 198 vol = 66.802
core size 120 of 198 vol = 62.901
core size 119 of 198 vol = 59.152
core size 118 of 198 vol = 55.75
core size 117 of 198 vol = 51.832
core size 116 of 198 vol = 48.3
core size 115 of 198 vol = 44.927
core size 114 of 198 vol = 42.418
core size 113 of 198 vol = 39.425
core size 112 of 198 vol = 37.381
core size 111 of 198 vol = 33.06
core size 110 of 198 vol = 28.153
core size 109 of 198 vol = 25.33
core size 108 of 198 vol = 22.509
core size 107 of 198 vol = 20.695
core size 106 of 198 vol = 18.754
core size 105 of 198 vol = 17.757
core size 104 of 198 vol = 16.712
core size 103 of 198 vol = 15.44
core size 102 of 198 vol = 14.745
core size 101 of 198 vol = 14.758
core size 100 of 198 vol = 13.11
core size 99 of 198 vol = 11.018
core size 98 of 198 vol = 8.967
core size 97 of 198 vol = 7.643
core size 96 of 198 vol = 6.326
core size 95 of 198 vol = 5.37
core size 94 of 198 vol = 4.312
core size 93 of 198 vol = 3.391
core size 92 of 198 vol = 2.697
core size 91 of 198 vol = 1.911
core size 90 of 198 vol = 1.577
core size 89 of 198 vol = 1.144
core size 88 of 198 vol = 0.826
core size 87 of 198 vol = 0.594
core size 86 of 198 vol = 0.494
FINISHED: Min vol ( 0.5 ) reached
core.inds <- print(core, vol=0.5)
# 87 positions (cumulative volume <= 0.5 Angstrom^3)
start end length
1 8 50 43
2 52 95 44
xyz <- pdbfit(pdbs, core.inds, outpath="corefit_structures")
rf <- rmsf(xyz)
plotb3(rf, sse=pdb)
abline(v=100, col="gray", ylab="RMSF")