Kavi's BGGN 213 Portfolio

Class work for bioinformatics class
Class 17
Kavi (PID: A69046927)
ENSEMBL/OMIM Calculations
library(dplyr)
Attaching package: 'dplyr'
The following objects are masked from 'package:stats':
filter, lag
The following objects are masked from 'package:base':
intersect, setdiff, setequal, union
read.csv("MXL_genotypes.csv") -> MXL_genotypes
table(MXL_genotypes)
, , Population.s. = ALL, AMR, MXL, Father = -, Mother = -
Genotype..forward.strand.
Sample..Male.Female.Unknown. A|A A|G G|A G|G
NA19648 (F) 1 0 0 0
NA19649 (M) 0 0 0 1
NA19651 (F) 1 0 0 0
NA19652 (M) 0 0 0 1
NA19654 (F) 0 0 0 1
NA19655 (M) 0 1 0 0
NA19657 (F) 0 1 0 0
NA19658 (M) 1 0 0 0
NA19661 (M) 0 1 0 0
NA19663 (F) 1 0 0 0
NA19664 (M) 0 0 1 0
NA19669 (F) 1 0 0 0
NA19670 (M) 1 0 0 0
NA19676 (M) 0 0 0 1
NA19678 (F) 1 0 0 0
NA19679 (M) 0 1 0 0
NA19681 (F) 0 1 0 0
NA19682 (M) 0 1 0 0
NA19684 (F) 0 1 0 0
NA19716 (F) 0 0 1 0
NA19717 (M) 0 1 0 0
NA19719 (F) 0 0 0 1
NA19720 (M) 0 0 0 1
NA19722 (F) 0 0 1 0
NA19723 (M) 0 0 0 1
NA19725 (F) 0 1 0 0
NA19726 (M) 1 0 0 0
NA19728 (F) 1 0 0 0
NA19729 (M) 0 1 0 0
NA19731 (F) 1 0 0 0
NA19732 (M) 0 1 0 0
NA19734 (F) 0 0 1 0
NA19735 (M) 0 0 0 1
NA19740 (F) 1 0 0 0
NA19741 (M) 1 0 0 0
NA19746 (F) 1 0 0 0
NA19747 (M) 0 0 1 0
NA19749 (F) 0 1 0 0
NA19750 (M) 0 1 0 0
NA19752 (F) 0 1 0 0
NA19755 (F) 1 0 0 0
NA19756 (M) 0 0 1 0
NA19758 (F) 0 1 0 0
NA19759 (M) 0 0 1 0
NA19761 (F) 0 0 1 0
NA19762 (M) 1 0 0 0
NA19764 (F) 1 0 0 0
NA19770 (F) 0 1 0 0
NA19771 (M) 1 0 0 0
NA19773 (F) 1 0 0 0
NA19774 (M) 0 1 0 0
NA19776 (F) 0 1 0 0
NA19777 (M) 1 0 0 0
NA19779 (F) 0 0 1 0
NA19780 (M) 1 0 0 0
NA19782 (F) 0 0 1 0
NA19783 (M) 0 1 0 0
NA19785 (F) 1 0 0 0
NA19786 (M) 0 0 1 0
NA19788 (F) 0 1 0 0
NA19789 (M) 0 0 0 1
NA19792 (M) 1 0 0 0
NA19794 (F) 0 0 1 0
NA19795 (M) 0 1 0 0
sum(MXL_genotypes$Genotype..forward.strand.=="G|G") -> homoz_numb
all_numb = count(MXL_genotypes)
homoz_prop <- (homoz_numb/all_numb) * 100
homoz_prop
n
1 14.0625
DESeq Analysis
genereads <- read.table("genereads.txt", header=TRUE, row.names=1)
summary(genereads)
sample geno exp
Length:462 Length:462 Min. : 6.675
Class :character Class :character 1st Qu.:20.004
Mode :character Mode :character Median :25.116
Mean :25.640
3rd Qu.:30.779
Max. :51.518
aa <- sum(genereads$geno == "A/A")
ag <- sum(genereads$geno == "A/G")
gg <- sum(genereads$geno == "G/G")
Q13 Sample Size
There are 108 individuals with A/A genotype, 233 individuals with
A/G genotype, and 121 individuals with G/G genotype.
Median expression provided in boxplot for Q14.
Q14 Box Plot
library(ggplot2)
ggplot(genereads) +
aes(x=geno,y=exp) +
theme_classic() +
geom_boxplot() +
stat_summary(
fun = median,
geom = "text",
aes(label = round(..y.., 2)),
vjust = -0.5,
color = "red"
) +
labs(x="Genotype", y="Expression Level") +
geom_jitter(alpha=0.2, width = 0.2)
Warning: The dot-dot notation (`..y..`) was deprecated in ggplot2 3.4.0.
ℹ Please use `after_stat(y)` instead.
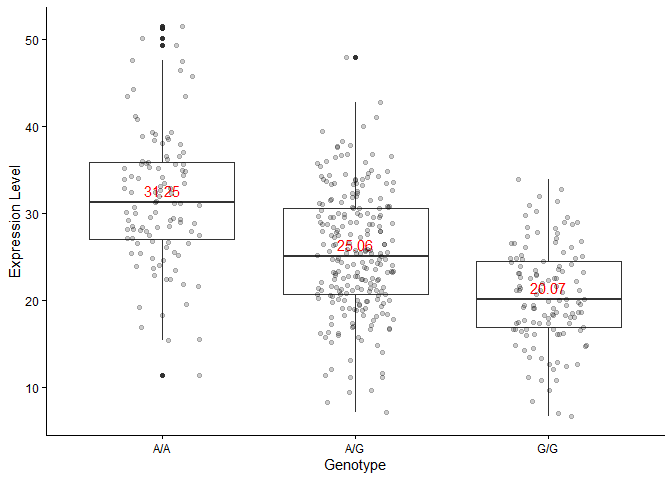
Response to Q14
From the boxplot, I can infer that the more copies there are of the SNP allele variant, the lower the expression of ORMDL3. The allele inhibits proper expression of ORMDL3.